Difference between revisions of "HAEM5:Myelodysplastic neoplasm with low blasts and 5q deletion"
| [unchecked revision] | [checked revision] |
Bailey.Glen (talk | contribs) |
Bailey.Glen (talk | contribs) |
||
| (4 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
{{DISPLAYTITLE:Myelodysplastic neoplasm with low blasts and 5q deletion}} | {{DISPLAYTITLE:Myelodysplastic neoplasm with low blasts and 5q deletion}} | ||
| − | [[HAEM5:Table_of_Contents|Haematolymphoid Tumours (5th ed.)]] | + | [[HAEM5:Table_of_Contents|Haematolymphoid Tumours (WHO Classification, 5th ed.)]] |
{{Under Construction}} | {{Under Construction}} | ||
| − | <blockquote class='blockedit'>{{Box-round|title= | + | <blockquote class='blockedit'>{{Box-round|title=Content Update To WHO 5th Edition Classification Is In Process; Content Below is Based on WHO 4th Edition Classification|This page was converted to the new template on 2023-12-07. The original page can be found at [[HAEM4:Myelodysplastic Syndrome (MDS) with Isolated del(5q)]]. |
}}</blockquote> | }}</blockquote> | ||
| + | |||
| + | <span style="color:#0070C0">(General Instructions – The main focus of these pages is the clinically significant genetic alterations in each disease type. Use [https://www.genenames.org/ <u>HUGO-approved gene names and symbols</u>] (italicized when appropriate), [https://varnomen.hgvs.org/ HGVS-based nomenclature for variants], as well as generic names of drugs and testing platforms or assays if applicable. Please complete tables whenever possible and do not delete them (add N/A if not applicable in the table and delete the examples); to add (or move) a row or column to a table, click within the table and select the > symbol that appears to be given options. Please do not delete or alter the section headings. The use of bullet points alongside short blocks of text rather than only large paragraphs is encouraged. Additional instructions below in italicized blue text should not be included in the final page content. Please also see </span><u>[[Author_Instructions]]</u><span style="color:#0070C0"> and [[Frequently Asked Questions (FAQs)|<u>FAQs</u>]] as well as contact your [[Leadership|<u>Associate Editor</u>]] or [mailto:CCGA@cancergenomics.org <u>Technical Support</u>])</span> | ||
| + | |||
==Primary Author(s)*== | ==Primary Author(s)*== | ||
| Line 14: | Line 17: | ||
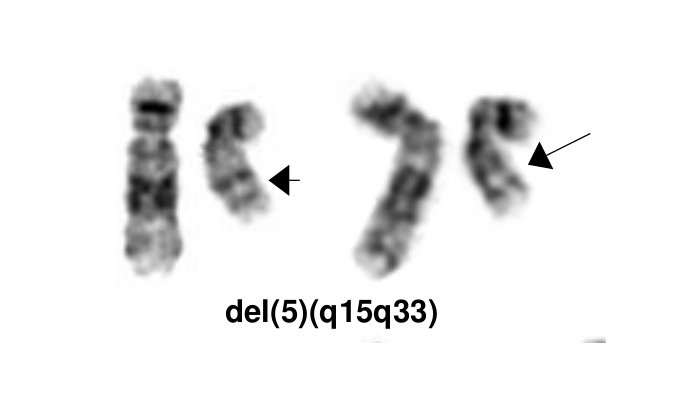
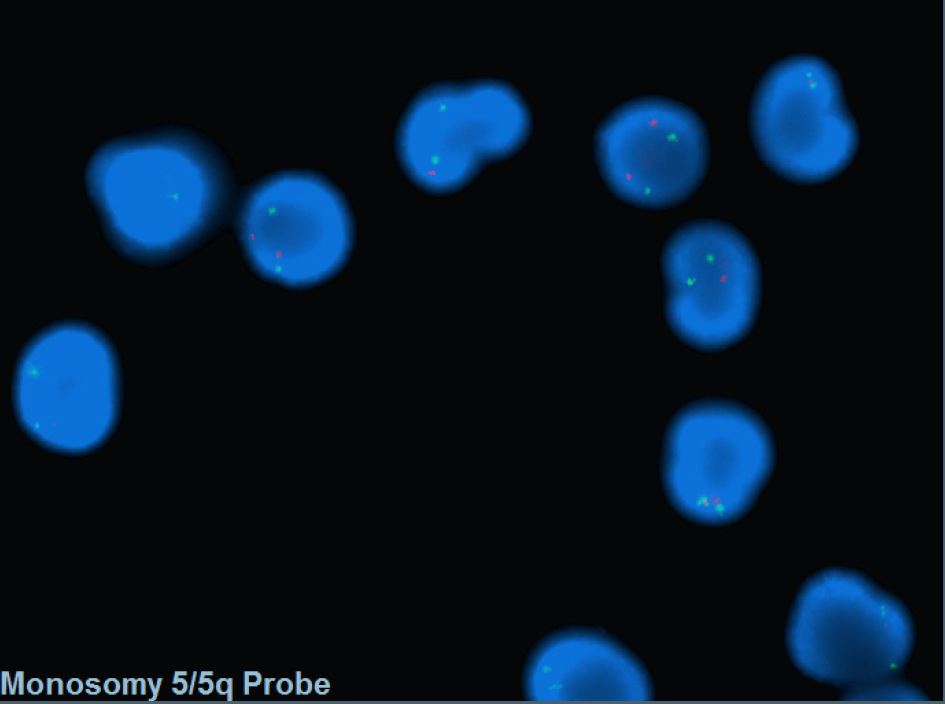
[[File:del(5)(q15q33).png|del(5)(q15q33)|322x322px]] | [[File:del(5)(q15q33).png|del(5)(q15q33)|322x322px]] | ||
| − | == | + | ==WHO Classification of Disease== |
| − | + | {| class="wikitable" | |
| − | + | !Structure | |
| − | + | !Disease | |
| − | + | |- | |
| − | Myelodysplastic | + | |Book |
| + | |Haematolymphoid Tumours (5th ed.) | ||
| + | |- | ||
| + | |Category | ||
| + | |Myeloid proliferations and neoplasms | ||
| + | |- | ||
| + | |Family | ||
| + | |Myelodysplastic neoplasms | ||
| + | |- | ||
| + | |Type | ||
| + | |Myelodysplastic neoplasms, with defining genetic abnormalities | ||
| + | |- | ||
| + | |Subtype(s) | ||
| + | |Myelodysplastic neoplasm with low blasts and 5q deletion | ||
| + | |} | ||
==Definition / Description of Disease== | ==Definition / Description of Disease== | ||
| − | MDS with isolated del(5q) is a type of [[Myelodysplastic Syndromes (MDS)|MDS]] with defining cytogenetic abnormality of del(5q). The World Health Organization named it as isolated del 5q, but occasionally the deletion can occur with an additional cytogenetic abnormality other than monosomy 7 or del(7q). The 5q- syndrome was first described by Van den Berghe et al as a distinct type of MDS featured with macrocytic anemia, hypolobulated megakaryocytes, a normal or increased platelet count <ref name=":0" />. Deletion of 5q is the most common recurrent cytogenetic abnormality in myeloid neoplasm and it was commonly seen in 10-15% of patients with MDS<ref>{{Cite journal|last=Hosono|first=Naoko|last2=Makishima|first2=Hideki|last3=Mahfouz|first3=Reda|last4=Przychodzen|first4=Bartlomiej|last5=Yoshida|first5=Kenichi|last6=Jerez|first6=Andres|last7=LaFramboise|first7=Thomas|last8=Polprasert|first8=Chantana|last9=Clemente|first9=Michael J|date=2017|title=Recurrent genetic defects on chromosome 5q in myeloid neoplasms|url=https://www.oncotarget.com/lookup/doi/10.18632/oncotarget.14130|journal=Oncotarget|language=en|volume=8|issue=4|pages=6483–6495|doi=10.18632/oncotarget.14130|issn=1949-2553|pmc=PMC5351647|pmid=28031539}}</ref>. A ~1.5 Mb common deleted region (CDR) at 5q32-q33 was identified in 5q- syndrome patients and was associated with good prognosis <ref>{{Cite journal|last=Boultwood|first=J.|last2=Fidler|first2=C.|last3=Lewis|first3=S.|last4=Kelly|first4=S.|last5=Sheridan|first5=H.|last6=Littlewood|first6=T.J.|last7=Buckle|first7=V.J.|last8=Wainscoat|first8=J.S.|date=1994|title=Molecular Mapping of Uncharacteristically Small 5q Deletions in Two Patients with the 5q- Syndrome: Delineation of the Critical Region on 5q and Identification of a 5q- Breakpoint|url=https://linkinghub.elsevier.com/retrieve/pii/S0888754384710901|journal=Genomics|language=en|volume=19|issue=3|pages=425–432|doi=10.1006/geno.1994.1090}}</ref>. This disease has a good respond to lenalidomide treatment (See Clinical Significance). | + | MDS with isolated del(5q) is a type of [[HAEM4:Myelodysplastic Syndromes (MDS)|MDS]] with defining cytogenetic abnormality of del(5q). The World Health Organization named it as isolated del 5q, but occasionally the deletion can occur with an additional cytogenetic abnormality other than monosomy 7 or del(7q). The 5q- syndrome was first described by Van den Berghe et al as a distinct type of MDS featured with macrocytic anemia, hypolobulated megakaryocytes, a normal or increased platelet count <ref name=":0" />. Deletion of 5q is the most common recurrent cytogenetic abnormality in myeloid neoplasm and it was commonly seen in 10-15% of patients with MDS<ref>{{Cite journal|last=Hosono|first=Naoko|last2=Makishima|first2=Hideki|last3=Mahfouz|first3=Reda|last4=Przychodzen|first4=Bartlomiej|last5=Yoshida|first5=Kenichi|last6=Jerez|first6=Andres|last7=LaFramboise|first7=Thomas|last8=Polprasert|first8=Chantana|last9=Clemente|first9=Michael J|date=2017|title=Recurrent genetic defects on chromosome 5q in myeloid neoplasms|url=https://www.oncotarget.com/lookup/doi/10.18632/oncotarget.14130|journal=Oncotarget|language=en|volume=8|issue=4|pages=6483–6495|doi=10.18632/oncotarget.14130|issn=1949-2553|pmc=PMC5351647|pmid=28031539}}</ref>. A ~1.5 Mb common deleted region (CDR) at 5q32-q33 was identified in 5q- syndrome patients and was associated with good prognosis <ref>{{Cite journal|last=Boultwood|first=J.|last2=Fidler|first2=C.|last3=Lewis|first3=S.|last4=Kelly|first4=S.|last5=Sheridan|first5=H.|last6=Littlewood|first6=T.J.|last7=Buckle|first7=V.J.|last8=Wainscoat|first8=J.S.|date=1994|title=Molecular Mapping of Uncharacteristically Small 5q Deletions in Two Patients with the 5q- Syndrome: Delineation of the Critical Region on 5q and Identification of a 5q- Breakpoint|url=https://linkinghub.elsevier.com/retrieve/pii/S0888754384710901|journal=Genomics|language=en|volume=19|issue=3|pages=425–432|doi=10.1006/geno.1994.1090}}</ref>. This disease has a good respond to lenalidomide treatment (See Clinical Significance). |
==Synonyms / Terminology== | ==Synonyms / Terminology== | ||
| Line 37: | Line 54: | ||
==Clinical Features== | ==Clinical Features== | ||
| − | Put your text here and fill in the table <span style="color:#0070C0">(''Instruction: Can include references in the table'') </span> | + | Put your text here and fill in the table <span style="color:#0070C0">(''Instruction: Can include references in the table. Do not delete table.'') </span> |
{| class="wikitable" | {| class="wikitable" | ||
|'''Signs and Symptoms''' | |'''Signs and Symptoms''' | ||
| − | |EXAMPLE Asymptomatic (incidental finding on complete blood counts) | + | |<span class="blue-text">EXAMPLE:</span> Asymptomatic (incidental finding on complete blood counts) |
| − | EXAMPLE B-symptoms (weight loss, fever, night sweats) | + | <span class="blue-text">EXAMPLE:</span> B-symptoms (weight loss, fever, night sweats) |
| − | EXAMPLE Fatigue | + | <span class="blue-text">EXAMPLE:</span> Fatigue |
| − | EXAMPLE Lymphadenopathy (uncommon) | + | <span class="blue-text">EXAMPLE:</span> Lymphadenopathy (uncommon) |
|- | |- | ||
|'''Laboratory Findings''' | |'''Laboratory Findings''' | ||
| − | |EXAMPLE Cytopenias | + | |<span class="blue-text">EXAMPLE:</span> Cytopenias |
| − | EXAMPLE Lymphocytosis (low level) | + | <span class="blue-text">EXAMPLE:</span> Lymphocytosis (low level) |
|} | |} | ||
| Line 74: | Line 91: | ||
==Immunophenotype== | ==Immunophenotype== | ||
| − | Put your text here and fill in the table <span style="color:#0070C0">(''Instruction: Can include references in the table'') </span> | + | Put your text here and fill in the table <span style="color:#0070C0">(''Instruction: Can include references in the table. Do not delete table.'') </span> |
{| class="wikitable sortable" | {| class="wikitable sortable" | ||
| Line 80: | Line 97: | ||
!Finding!!Marker | !Finding!!Marker | ||
|- | |- | ||
| − | |Positive (universal)||EXAMPLE CD1 | + | |Positive (universal)||<span class="blue-text">EXAMPLE:</span> CD1 |
|- | |- | ||
| − | |Positive (subset)||EXAMPLE CD2 | + | |Positive (subset)||<span class="blue-text">EXAMPLE:</span> CD2 |
|- | |- | ||
| − | |Negative (universal)||EXAMPLE CD3 | + | |Negative (universal)||<span class="blue-text">EXAMPLE:</span> CD3 |
|- | |- | ||
| − | |Negative (subset)||EXAMPLE CD4 | + | |Negative (subset)||<span class="blue-text">EXAMPLE:</span> CD4 |
|} | |} | ||
| Line 107: | Line 124: | ||
!Notes | !Notes | ||
|- | |- | ||
| − | |EXAMPLE t(9;22)(q34;q11.2)||EXAMPLE 3'ABL1 / 5'BCR||EXAMPLE der(22)||EXAMPLE 20% (COSMIC) | + | |<span class="blue-text">EXAMPLE:</span> t(9;22)(q34;q11.2)||<span class="blue-text">EXAMPLE:</span> 3'ABL1 / 5'BCR||<span class="blue-text">EXAMPLE:</span> der(22)||<span class="blue-text">EXAMPLE:</span> 20% (COSMIC) |
| − | EXAMPLE 30% (add reference) | + | <span class="blue-text">EXAMPLE:</span> 30% (add reference) |
|Yes | |Yes | ||
|No | |No | ||
|Yes | |Yes | ||
| − | |EXAMPLE | + | |<span class="blue-text">EXAMPLE:</span> |
The t(9;22) is diagnostic of CML in the appropriate morphology and clinical context (add reference). This fusion is responsive to targeted therapy such as Imatinib (Gleevec) (add reference). | The t(9;22) is diagnostic of CML in the appropriate morphology and clinical context (add reference). This fusion is responsive to targeted therapy such as Imatinib (Gleevec) (add reference). | ||
| Line 126: | Line 143: | ||
!Chromosomal Rearrangement!!Genes in Fusion (5’ or 3’ Segments)!!Pathogenic Derivative!!Prevalence | !Chromosomal Rearrangement!!Genes in Fusion (5’ or 3’ Segments)!!Pathogenic Derivative!!Prevalence | ||
|- | |- | ||
| − | |EXAMPLE t(9;22)(q34;q11.2)||EXAMPLE 3'ABL1 / 5'BCR||EXAMPLE der(22)||EXAMPLE 5% | + | |<span class="blue-text">EXAMPLE:</span> t(9;22)(q34;q11.2)||<span class="blue-text">EXAMPLE:</span> 3'ABL1 / 5'BCR||<span class="blue-text">EXAMPLE:</span> der(22)||<span class="blue-text">EXAMPLE:</span> 5% |
|- | |- | ||
| − | |EXAMPLE t(8;21)(q22;q22)||EXAMPLE 5'RUNX1 / 3'RUNXT1||EXAMPLE der(8)||EXAMPLE 5% | + | |<span class="blue-text">EXAMPLE:</span> t(8;21)(q22;q22)||<span class="blue-text">EXAMPLE:</span> 5'RUNX1 / 3'RUNXT1||<span class="blue-text">EXAMPLE:</span> der(8)||<span class="blue-text">EXAMPLE:</span> 5% |
|} | |} | ||
| Line 161: | Line 178: | ||
==Individual Region Genomic Gain / Loss / LOH== | ==Individual Region Genomic Gain / Loss / LOH== | ||
| − | Put your text here and fill in the table <span style="color:#0070C0">(''Instructions: Includes aberrations not involving gene fusions. Can include references in the table. Can refer to CGC workgroup tables as linked on the homepage if applicable.'') </span> | + | Put your text here and fill in the table <span style="color:#0070C0">(''Instructions: Includes aberrations not involving gene fusions. Can include references in the table. Can refer to CGC workgroup tables as linked on the homepage if applicable. Do not delete table.'') </span> |
{| class="wikitable sortable" | {| class="wikitable sortable" | ||
| Line 171: | Line 188: | ||
!Notes | !Notes | ||
|- | |- | ||
| − | |EXAMPLE | + | |<span class="blue-text">EXAMPLE:</span> |
7 | 7 | ||
| − | |EXAMPLE Loss | + | |<span class="blue-text">EXAMPLE:</span> Loss |
| − | |EXAMPLE | + | |<span class="blue-text">EXAMPLE:</span> |
chr7:1- 159,335,973 [hg38] | chr7:1- 159,335,973 [hg38] | ||
| − | |EXAMPLE | + | |<span class="blue-text">EXAMPLE:</span> |
chr7 | chr7 | ||
| Line 184: | Line 201: | ||
|Yes | |Yes | ||
|No | |No | ||
| − | |EXAMPLE | + | |<span class="blue-text">EXAMPLE:</span> |
Presence of monosomy 7 (or 7q deletion) is sufficient for a diagnosis of AML with MDS-related changes when there is ≥20% blasts and no prior therapy (add reference). Monosomy 7/7q deletion is associated with a poor prognosis in AML (add reference). | Presence of monosomy 7 (or 7q deletion) is sufficient for a diagnosis of AML with MDS-related changes when there is ≥20% blasts and no prior therapy (add reference). Monosomy 7/7q deletion is associated with a poor prognosis in AML (add reference). | ||
|- | |- | ||
| − | |EXAMPLE | + | |<span class="blue-text">EXAMPLE:</span> |
8 | 8 | ||
| − | |EXAMPLE Gain | + | |<span class="blue-text">EXAMPLE:</span> Gain |
| − | |EXAMPLE | + | |<span class="blue-text">EXAMPLE:</span> |
chr8:1-145,138,636 [hg38] | chr8:1-145,138,636 [hg38] | ||
| − | |EXAMPLE | + | |<span class="blue-text">EXAMPLE:</span> |
chr8 | chr8 | ||
| Line 201: | Line 218: | ||
|No | |No | ||
|No | |No | ||
| − | |EXAMPLE | + | |<span class="blue-text">EXAMPLE:</span> |
Common recurrent secondary finding for t(8;21) (add reference). | Common recurrent secondary finding for t(8;21) (add reference). | ||
| Line 212: | Line 229: | ||
!Chromosome Number!!Gain/Loss/Amp/LOH!!Region | !Chromosome Number!!Gain/Loss/Amp/LOH!!Region | ||
|- | |- | ||
| − | |EXAMPLE 8||EXAMPLE Gain||EXAMPLE chr8:0-1000000 | + | |<span class="blue-text">EXAMPLE:</span> 8||<span class="blue-text">EXAMPLE:</span> Gain||<span class="blue-text">EXAMPLE:</span> chr8:0-1000000 |
|- | |- | ||
| − | |EXAMPLE 7||EXAMPLE Loss||EXAMPLE chr7:0-1000000 | + | |<span class="blue-text">EXAMPLE:</span> 7||<span class="blue-text">EXAMPLE:</span> Loss||<span class="blue-text">EXAMPLE:</span> chr7:0-1000000 |
|} | |} | ||
| Line 220: | Line 237: | ||
==Characteristic Chromosomal Patterns== | ==Characteristic Chromosomal Patterns== | ||
| − | Put your text here <span style="color:#0070C0">(''EXAMPLE PATTERNS: hyperdiploid; gain of odd number chromosomes including typically chromosome 1, 3, 5, 7, 11, and 17; co-deletion of 1p and 19q; complex karyotypes without characteristic genetic findings; chromothripsis'')</span> | + | Put your text here <span style="color:#0070C0">(''EXAMPLE PATTERNS: hyperdiploid; gain of odd number chromosomes including typically chromosome 1, 3, 5, 7, 11, and 17; co-deletion of 1p and 19q; complex karyotypes without characteristic genetic findings; chromothripsis. Do not delete table.'')</span> |
{| class="wikitable sortable" | {| class="wikitable sortable" | ||
| Line 230: | Line 247: | ||
!Notes | !Notes | ||
|- | |- | ||
| − | |EXAMPLE | + | |<span class="blue-text">EXAMPLE:</span> |
Co-deletion of 1p and 18q | Co-deletion of 1p and 18q | ||
| Line 236: | Line 253: | ||
|No | |No | ||
|No | |No | ||
| − | |EXAMPLE: | + | |<span class="blue-text">EXAMPLE:</span> |
See chromosomal rearrangements table as this pattern is due to an unbalanced derivative translocation associated with oligodendroglioma (add reference). | See chromosomal rearrangements table as this pattern is due to an unbalanced derivative translocation associated with oligodendroglioma (add reference). | ||
| Line 247: | Line 264: | ||
==Gene Mutations (SNV / INDEL)== | ==Gene Mutations (SNV / INDEL)== | ||
| − | Put your text here and fill in the table <span style="color:#0070C0">(''Instructions: This table is not meant to be an exhaustive list; please include only genes/alterations that are recurrent and common as well either disease defining and/or clinically significant. Can include references in the table. For clinical significance, denote associations with FDA-approved therapy (not an extensive list of applicable drugs) and NCCN or other national guidelines if applicable | + | Put your text here and fill in the table <span style="color:#0070C0">(''Instructions: This table is not meant to be an exhaustive list; please include only genes/alterations that are recurrent and common as well as either disease defining and/or clinically significant. Can include references in the table. For clinical significance, denote associations with FDA-approved therapy (not an extensive list of applicable drugs) and NCCN or other national guidelines if applicable. Can also refer to CGC workgroup tables as linked on the homepage if applicable as well as any high impact papers or reviews of gene mutations in this entity. Do not delete table.'') </span> |
{| class="wikitable sortable" | {| class="wikitable sortable" | ||
| Line 257: | Line 274: | ||
!Notes | !Notes | ||
|- | |- | ||
| − | |EXAMPLE: TP53; Variable LOF mutations | + | |<span class="blue-text">EXAMPLE:</span> TP53; Variable LOF mutations |
| − | EXAMPLE: | + | <span class="blue-text">EXAMPLE:</span> |
EGFR; Exon 20 mutations | EGFR; Exon 20 mutations | ||
| − | EXAMPLE: BRAF; Activating mutations | + | <span class="blue-text">EXAMPLE:</span> BRAF; Activating mutations |
| − | |EXAMPLE: TSG | + | |<span class="blue-text">EXAMPLE:</span> TSG |
| − | |EXAMPLE: 20% (COSMIC) | + | |<span class="blue-text">EXAMPLE:</span> 20% (COSMIC) |
| − | EXAMPLE: 30% (add Reference) | + | <span class="blue-text">EXAMPLE:</span> 30% (add Reference) |
| − | |EXAMPLE: IDH1 R123H | + | |<span class="blue-text">EXAMPLE:</span> IDH1 R123H |
| − | |EXAMPLE: EGFR amplification | + | |<span class="blue-text">EXAMPLE:</span> EGFR amplification |
| | | | ||
| | | | ||
| | | | ||
| − | |EXAMPLE: Excludes hairy cell leukemia (HCL) (add reference). | + | |<span class="blue-text">EXAMPLE:</span> Excludes hairy cell leukemia (HCL) (add reference). |
<br /> | <br /> | ||
|} | |} | ||
| Line 287: | Line 304: | ||
!Gene!!Mutation!!Oncogene/Tumor Suppressor/Other!!Presumed Mechanism (LOF/GOF/Other; Driver/Passenger)!!Prevalence (COSMIC/TCGA/Other) | !Gene!!Mutation!!Oncogene/Tumor Suppressor/Other!!Presumed Mechanism (LOF/GOF/Other; Driver/Passenger)!!Prevalence (COSMIC/TCGA/Other) | ||
|- | |- | ||
| − | |EXAMPLE TP53||EXAMPLE R273H||EXAMPLE Tumor Suppressor||EXAMPLE LOF||EXAMPLE 20% | + | |<span class="blue-text">EXAMPLE:</span> TP53||<span class="blue-text">EXAMPLE:</span> R273H||<span class="blue-text">EXAMPLE:</span> Tumor Suppressor||<span class="blue-text">EXAMPLE:</span> LOF||<span class="blue-text">EXAMPLE:</span> 20% |
|} | |} | ||
| Line 309: | Line 326: | ||
==Genes and Main Pathways Involved== | ==Genes and Main Pathways Involved== | ||
| − | Put your text here and fill in the table <span style="color:#0070C0">(''Instructions: Can include references in the table.'')</span> | + | Put your text here and fill in the table <span style="color:#0070C0">(''Instructions: Can include references in the table. Do not delete table.'')</span> |
{| class="wikitable sortable" | {| class="wikitable sortable" | ||
|- | |- | ||
!Gene; Genetic Alteration!!Pathway!!Pathophysiologic Outcome | !Gene; Genetic Alteration!!Pathway!!Pathophysiologic Outcome | ||
|- | |- | ||
| − | |EXAMPLE: BRAF and MAP2K1; Activating mutations | + | |<span class="blue-text">EXAMPLE:</span> BRAF and MAP2K1; Activating mutations |
| − | |EXAMPLE: MAPK signaling | + | |<span class="blue-text">EXAMPLE:</span> MAPK signaling |
| − | |EXAMPLE: Increased cell growth and proliferation | + | |<span class="blue-text">EXAMPLE:</span> Increased cell growth and proliferation |
|- | |- | ||
| − | |EXAMPLE: CDKN2A; Inactivating mutations | + | |<span class="blue-text">EXAMPLE:</span> CDKN2A; Inactivating mutations |
| − | |EXAMPLE: Cell cycle regulation | + | |<span class="blue-text">EXAMPLE:</span> Cell cycle regulation |
| − | |EXAMPLE: Unregulated cell division | + | |<span class="blue-text">EXAMPLE:</span> Unregulated cell division |
|- | |- | ||
| − | |EXAMPLE: KMT2C and ARID1A; Inactivating mutations | + | |<span class="blue-text">EXAMPLE:</span> KMT2C and ARID1A; Inactivating mutations |
| − | |EXAMPLE: Histone modification, chromatin remodeling | + | |<span class="blue-text">EXAMPLE:</span> Histone modification, chromatin remodeling |
| − | |EXAMPLE: Abnormal gene expression program | + | |<span class="blue-text">EXAMPLE:</span> Abnormal gene expression program |
|} | |} | ||
Latest revision as of 17:16, 6 September 2024
Haematolymphoid Tumours (WHO Classification, 5th ed.)
| This page is under construction |
editContent Update To WHO 5th Edition Classification Is In Process; Content Below is Based on WHO 4th Edition ClassificationThis page was converted to the new template on 2023-12-07. The original page can be found at HAEM4:Myelodysplastic Syndrome (MDS) with Isolated del(5q).
(General Instructions – The main focus of these pages is the clinically significant genetic alterations in each disease type. Use HUGO-approved gene names and symbols (italicized when appropriate), HGVS-based nomenclature for variants, as well as generic names of drugs and testing platforms or assays if applicable. Please complete tables whenever possible and do not delete them (add N/A if not applicable in the table and delete the examples); to add (or move) a row or column to a table, click within the table and select the > symbol that appears to be given options. Please do not delete or alter the section headings. The use of bullet points alongside short blocks of text rather than only large paragraphs is encouraged. Additional instructions below in italicized blue text should not be included in the final page content. Please also see Author_Instructions and FAQs as well as contact your Associate Editor or Technical Support)
Primary Author(s)*
Xiaolin Hu, Ph.D; Teresa Smolarek, Ph.D, FACMG
WHO Classification of Disease
| Structure | Disease |
|---|---|
| Book | Haematolymphoid Tumours (5th ed.) |
| Category | Myeloid proliferations and neoplasms |
| Family | Myelodysplastic neoplasms |
| Type | Myelodysplastic neoplasms, with defining genetic abnormalities |
| Subtype(s) | Myelodysplastic neoplasm with low blasts and 5q deletion |
Definition / Description of Disease
MDS with isolated del(5q) is a type of MDS with defining cytogenetic abnormality of del(5q). The World Health Organization named it as isolated del 5q, but occasionally the deletion can occur with an additional cytogenetic abnormality other than monosomy 7 or del(7q). The 5q- syndrome was first described by Van den Berghe et al as a distinct type of MDS featured with macrocytic anemia, hypolobulated megakaryocytes, a normal or increased platelet count [1]. Deletion of 5q is the most common recurrent cytogenetic abnormality in myeloid neoplasm and it was commonly seen in 10-15% of patients with MDS[2]. A ~1.5 Mb common deleted region (CDR) at 5q32-q33 was identified in 5q- syndrome patients and was associated with good prognosis [3]. This disease has a good respond to lenalidomide treatment (See Clinical Significance).
Synonyms / Terminology
Myelodysplastic syndromes with 5q deletion; 5q minus syndrome
Epidemiology / Prevalence
- Median age 67 years [4]
- Women versus male about 2:1
Clinical Features
Put your text here and fill in the table (Instruction: Can include references in the table. Do not delete table.)
| Signs and Symptoms | EXAMPLE: Asymptomatic (incidental finding on complete blood counts)
EXAMPLE: B-symptoms (weight loss, fever, night sweats) EXAMPLE: Fatigue EXAMPLE: Lymphadenopathy (uncommon) |
| Laboratory Findings | EXAMPLE: Cytopenias
EXAMPLE: Lymphocytosis (low level) |
editv4:Clinical FeaturesThe content below was from the old template. Please incorporate above.
- Macrocytic anemia most common, usually severe and can be transfusion dependent.
- Thrombocytosis can be seen in 1/3 to 1/2 of cases, whereas thrombocytopenia or neutropenia are uncommon.
- Pancytopenia is rare. If pancytopenia with isolated 5q-, MDS with unclassifiable should be considered. [WHO]
- Good prognosis and low risk to progress to AML.
Sites of Involvement
Blood or bone marrow
Morphologic Features
On bone marrow examination, cellularity is usually normal or hypercellular. Erythroid hypoplasia may be seen. Blasts cells are < 5% in the bone marrow and < 1% in the peripheral blood. The most characteristic feature of isolated 5q- is the presence of micromegakaryocytes with hypolobated nuclei [insert pic micromegakaryocyte]. The erythroid dysplasia may be seen but not predominant. Granulocytic dysplasia is uncommon.
Hypolobated micromegakaryocytes are also seen in CML, which is characterized by leukocytosis, basophilia, and myeloid hyperplasia with leftward shift. Platelet count increase can be seen in essential thrombocythemia (ET), but ET has large megakaryocytes with hyperlobulated nuclei rather than microkaryocytes with hypolobulation in MDS with isolated del(5q).
Immunophenotype
Put your text here and fill in the table (Instruction: Can include references in the table. Do not delete table.)
| Finding | Marker |
|---|---|
| Positive (universal) | EXAMPLE: CD1 |
| Positive (subset) | EXAMPLE: CD2 |
| Negative (universal) | EXAMPLE: CD3 |
| Negative (subset) | EXAMPLE: CD4 |
editv4:ImmunophenotypeThe content below was from the old template. Please incorporate above.There is no distinct immunophenotypic profile specific for myelodysplastic syndrome (MDS) with isolated del(5q). Currently, morphologic evaluation remains the gold standard in diagnosis of MDS. Immunophenotyping provides supportive evidence to clarify the blasts nature and percentage [5].
Chromosomal Rearrangements (Gene Fusions)
Put your text here and fill in the table
| Chromosomal Rearrangement | Genes in Fusion (5’ or 3’ Segments) | Pathogenic Derivative | Prevalence | Diagnostic Significance (Yes, No or Unknown) | Prognostic Significance (Yes, No or Unknown) | Therapeutic Significance (Yes, No or Unknown) | Notes |
|---|---|---|---|---|---|---|---|
| EXAMPLE: t(9;22)(q34;q11.2) | EXAMPLE: 3'ABL1 / 5'BCR | EXAMPLE: der(22) | EXAMPLE: 20% (COSMIC)
EXAMPLE: 30% (add reference) |
Yes | No | Yes | EXAMPLE:
The t(9;22) is diagnostic of CML in the appropriate morphology and clinical context (add reference). This fusion is responsive to targeted therapy such as Imatinib (Gleevec) (add reference). |
editv4:Chromosomal Rearrangements (Gene Fusions)The content below was from the old template. Please incorporate above.NA
Chromosomal Rearrangement Genes in Fusion (5’ or 3’ Segments) Pathogenic Derivative Prevalence EXAMPLE: t(9;22)(q34;q11.2) EXAMPLE: 3'ABL1 / 5'BCR EXAMPLE: der(22) EXAMPLE: 5% EXAMPLE: t(8;21)(q22;q22) EXAMPLE: 5'RUNX1 / 3'RUNXT1 EXAMPLE: der(8) EXAMPLE: 5%
editv4:Clinical Significance (Diagnosis, Prognosis and Therapeutic Implications).Please incorporate this section into the relevant tables found in:
- Chromosomal Rearrangements (Gene Fusions)
- Individual Region Genomic Gain/Loss/LOH
- Characteristic Chromosomal Patterns
- Gene Mutations (SNV/INDEL)
Diagnosis:
- Cytopenia in 1 to 2 lineages. Cytopenias are defined as hemoglobin concentration < 10 g/dL, platelet count < 100 x 109/L and absolute neutrophil count < 1.8 x 109/L; mild degree of anemia (hemoglobin < 13 g/dL in men or < 12 g/dL in women) or thrombocytopenia (platelets < 150 x109/L) are allowed if defining cytogenetic abnormality is present. Thrombocytosis (platelet count ≥ 450 x 109/L) is allowed in MDS with isolated del(5q). PB monocytes must be < 1 x109/L.
- Dysplasia in 1 to 3 lineages. Micromegakaryocytes with hypolobation are characteristic dysplasia. Ring sideroblasts are not common.
- Blasts percentage: BM <5%, PB <1%, no Auer rods.
- Interstitial deletion of 5q alone or with an additional abnormality other than loss of chromosome 7 or deletion of 7q.
- Chromosome 7 loss or del(7q), more than one additional cytogenetic abnormality, excess blasts need to be excluded from the diagnosis.
Prognosis:
- According to IPSS‐R good‐risk group with low rates of leukemic transformation [6].
- The median survival of MDS with isolated del(5q) is 66 to 145 months.
- About less than 10% of cases progress to acute myeloid leukemia [7].
- Mutations of TP53 may function as an independent prognostic factor in associated with leukemic progression and resistant to lenalidomide[8][9].
Therapeutic Implications:
Lenalidomide is an analogue of thalidomide which functions as an immunomodulatory agent. Treatment with lenalidomide has shown great potential in reducing the abnormal clone as well as transfusion dependency irrespective of cytogenetic complexity [10][11]. FDA approved Lenalidomide on December 2005 for treating MDS with del(5q) with or without additional cytogenetic abnormalities.
Individual Region Genomic Gain / Loss / LOH
Put your text here and fill in the table (Instructions: Includes aberrations not involving gene fusions. Can include references in the table. Can refer to CGC workgroup tables as linked on the homepage if applicable. Do not delete table.)
| Chr # | Gain / Loss / Amp / LOH | Minimal Region Genomic Coordinates [Genome Build] | Minimal Region Cytoband | Diagnostic Significance (Yes, No or Unknown) | Prognostic Significance (Yes, No or Unknown) | Therapeutic Significance (Yes, No or Unknown) | Notes |
|---|---|---|---|---|---|---|---|
| EXAMPLE:
7 |
EXAMPLE: Loss | EXAMPLE:
chr7:1- 159,335,973 [hg38] |
EXAMPLE:
chr7 |
Yes | Yes | No | EXAMPLE:
Presence of monosomy 7 (or 7q deletion) is sufficient for a diagnosis of AML with MDS-related changes when there is ≥20% blasts and no prior therapy (add reference). Monosomy 7/7q deletion is associated with a poor prognosis in AML (add reference). |
| EXAMPLE:
8 |
EXAMPLE: Gain | EXAMPLE:
chr8:1-145,138,636 [hg38] |
EXAMPLE:
chr8 |
No | No | No | EXAMPLE:
Common recurrent secondary finding for t(8;21) (add reference). |
editv4:Genomic Gain/Loss/LOHThe content below was from the old template. Please incorporate above.
Chromosome Number Gain/Loss/Amp/LOH Region EXAMPLE: 8 EXAMPLE: Gain EXAMPLE: chr8:0-1000000 EXAMPLE: 7 EXAMPLE: Loss EXAMPLE: chr7:0-1000000
Characteristic Chromosomal Patterns
Put your text here (EXAMPLE PATTERNS: hyperdiploid; gain of odd number chromosomes including typically chromosome 1, 3, 5, 7, 11, and 17; co-deletion of 1p and 19q; complex karyotypes without characteristic genetic findings; chromothripsis. Do not delete table.)
| Chromosomal Pattern | Diagnostic Significance (Yes, No or Unknown) | Prognostic Significance (Yes, No or Unknown) | Therapeutic Significance (Yes, No or Unknown) | Notes |
|---|---|---|---|---|
| EXAMPLE:
Co-deletion of 1p and 18q |
Yes | No | No | EXAMPLE:
See chromosomal rearrangements table as this pattern is due to an unbalanced derivative translocation associated with oligodendroglioma (add reference). |
editv4:Characteristic Chromosomal Aberrations / PatternsThe content below was from the old template. Please incorporate above.The most characteristic cytogenetic abnormality is an interstitial deletion on the long arm of chromosome 5 or del(5q)[1] The break point is not fixed but the region between bands q31 and q33 is generally deleted. Several candidates genes were thought to contribute to the haploinsufficiency effect of the deleted region, including RPS14 [12], CSNK1A1 [13], miR-145 and miR-146a [14]. Identification of del(5q) has clinical significance because patients with this cytogenetic abnormality have a good prognosis and they respond well to lenalidomide treatment.
Gene Mutations (SNV / INDEL)
Put your text here and fill in the table (Instructions: This table is not meant to be an exhaustive list; please include only genes/alterations that are recurrent and common as well as either disease defining and/or clinically significant. Can include references in the table. For clinical significance, denote associations with FDA-approved therapy (not an extensive list of applicable drugs) and NCCN or other national guidelines if applicable. Can also refer to CGC workgroup tables as linked on the homepage if applicable as well as any high impact papers or reviews of gene mutations in this entity. Do not delete table.)
| Gene; Genetic Alteration | Presumed Mechanism (Tumor Suppressor Gene [TSG] / Oncogene / Other) | Prevalence (COSMIC / TCGA / Other) | Concomitant Mutations | Mutually Exclusive Mutations | Diagnostic Significance (Yes, No or Unknown) | Prognostic Significance (Yes, No or Unknown) | Therapeutic Significance (Yes, No or Unknown) | Notes |
|---|---|---|---|---|---|---|---|---|
| EXAMPLE: TP53; Variable LOF mutations
EXAMPLE: EGFR; Exon 20 mutations EXAMPLE: BRAF; Activating mutations |
EXAMPLE: TSG | EXAMPLE: 20% (COSMIC)
EXAMPLE: 30% (add Reference) |
EXAMPLE: IDH1 R123H | EXAMPLE: EGFR amplification | EXAMPLE: Excludes hairy cell leukemia (HCL) (add reference).
|
Note: A more extensive list of mutations can be found in cBioportal (https://www.cbioportal.org/), COSMIC (https://cancer.sanger.ac.uk/cosmic), ICGC (https://dcc.icgc.org/) and/or other databases. When applicable, gene-specific pages within the CCGA site directly link to pertinent external content.
editv4:Gene Mutations (SNV/INDEL)The content below was from the old template. Please incorporate above.Somatic mutations in JAK2 and MPL have been reported in a small subset patients with isolated del(5q), but these mutations seem not confer diagnostic or prognostic value [15]. A subset of cases could have SF3B1 mutations, which needs to be differentiated with MDS-RS.
Gene Mutation Oncogene/Tumor Suppressor/Other Presumed Mechanism (LOF/GOF/Other; Driver/Passenger) Prevalence (COSMIC/TCGA/Other) EXAMPLE: TP53 EXAMPLE: R273H EXAMPLE: Tumor Suppressor EXAMPLE: LOF EXAMPLE: 20% Other Mutations
Type Gene/Region/Other Concomitant Mutations JAK2 V617F, MPL W515L Secondary Mutations Mutually Exclusive
Epigenomic Alterations
Genes involved in epigenetic regulation are frequently mutated in MDS such as TET2, DNMT3A, IDH1, IDH2, AXSL1, and EZH2 [16]. These genes play a role in DNA methylation and chromatin modification as well as regulating gene expression.
Genes and Main Pathways Involved
Put your text here and fill in the table (Instructions: Can include references in the table. Do not delete table.)
| Gene; Genetic Alteration | Pathway | Pathophysiologic Outcome |
|---|---|---|
| EXAMPLE: BRAF and MAP2K1; Activating mutations | EXAMPLE: MAPK signaling | EXAMPLE: Increased cell growth and proliferation |
| EXAMPLE: CDKN2A; Inactivating mutations | EXAMPLE: Cell cycle regulation | EXAMPLE: Unregulated cell division |
| EXAMPLE: KMT2C and ARID1A; Inactivating mutations | EXAMPLE: Histone modification, chromatin remodeling | EXAMPLE: Abnormal gene expression program |
editv4:Genes and Main Pathways InvolvedThe content below was from the old template. Please incorporate above.Several candidate genes in the common deleted region of 5q have been reported to contribute to the molecular pathogenesis.
RPS14: Encodes for ribosomal subunit of 40S. Inactivation of RPS14 results in ineffective erythroid differentiation and increased apoptosis in a p53 dependent way [17].
CSNK1A1: Encodes for CK1α, a serine/threonine protein kinase activity. Mutations in CSNK1A1 lead to clonal expansion of hematopoietic stem cells through activation of β-catenin [13].
miR-145 and miR146α: These two micro RNA genes are thought to be associated with thrombocytosis and hypolobulated megakaryocytes [18].
Genetic Diagnostic Testing Methods
- Bone marrow morphology and peripheral blood test are standard in diagnosis of MDS.
- FISH probes targeting the CDR are commonly incorporated into MDS panel to assess -5/5q-.
- Conventional cytogenetics are also helpful in diagnosis.
Familial Forms
Put your text here (Instructions: Include associated hereditary conditions/syndromes that cause this entity or are caused by this entity.)
Additional Information
Put your text here
Links
Put your links here (use link icon at top of page)
References
(use the "Cite" icon at the top of the page) (Instructions: Add each reference into the text above by clicking on where you want to insert the reference, selecting the “Cite” icon at the top of the page, and using the “Automatic” tab option to search such as by PMID to select the reference to insert. The reference list in this section will be automatically generated and sorted. If a PMID is not available, such as for a book, please use the “Cite” icon, select “Manual” and then “Basic Form”, and include the entire reference.)
- ↑ 1.0 1.1 Van Den Berghe, Herman; et al. (1974). "Distinct haematological disorder with deletion of long arm of No. 5 chromosome". Nature. 251 (5474): 437–438. doi:10.1038/251437a0. ISSN 0028-0836.
- ↑ Hosono, Naoko; et al. (2017). "Recurrent genetic defects on chromosome 5q in myeloid neoplasms". Oncotarget. 8 (4): 6483–6495. doi:10.18632/oncotarget.14130. ISSN 1949-2553. PMC 5351647. PMID 28031539.CS1 maint: PMC format (link)
- ↑ Boultwood, J.; et al. (1994). "Molecular Mapping of Uncharacteristically Small 5q Deletions in Two Patients with the 5q- Syndrome: Delineation of the Critical Region on 5q and Identification of a 5q- Breakpoint". Genomics. 19 (3): 425–432. doi:10.1006/geno.1994.1090.
- ↑ Giagounidis, A.A.N.; et al. (2004). "Hematological Malignancies". Hematology. 9 (4): 271–277. doi:10.1080/10245330410001723824. ISSN 1607-8454.
- ↑ Zini, Gina (2017). "Diagnostics and Prognostication of Myelodysplastic Syndromes". Annals of Laboratory Medicine. 37 (6): 465. doi:10.3343/alm.2017.37.6.465. ISSN 2234-3806. PMC 5587818. PMID 28840983.CS1 maint: PMC format (link)
- ↑ Mallo, M; et al. (2011). "Impact of adjunct cytogenetic abnormalities for prognostic stratification in patients with myelodysplastic syndrome and deletion 5q". Leukemia. 25 (1): 110–120. doi:10.1038/leu.2010.231. ISSN 0887-6924.
- ↑ Boultwood, Jacqueline; et al. (2010). "Advances in the 5q− syndrome". Blood. 116 (26): 5803–5811. doi:10.1182/blood-2010-04-273771. ISSN 0006-4971.
- ↑ Jädersten, Martin; et al. (2011). "TP53 Mutations in Low-Risk Myelodysplastic Syndromes With del(5q) Predict Disease Progression". Journal of Clinical Oncology. 29 (15): 1971–1979. doi:10.1200/JCO.2010.31.8576. ISSN 0732-183X.
- ↑ Kulasekararaj, Austin G.; et al. (2013). "TP53 mutations in myelodysplastic syndrome are strongly correlated with aberrations of chromosome 5, and correlate with adverse prognosis". British Journal of Haematology. 160 (5): 660–672. doi:10.1111/bjh.12203.
- ↑ Giagounidis, A. A. N.; et al. (2005). "Prognosis of patients with del(5q) MDS and complex karyotype and the possible role of lenalidomide in this patient subgroup". Annals of Hematology. 84 (9): 569–571. doi:10.1007/s00277-005-1054-0. ISSN 0939-5555.
- ↑ List, Alan; et al. (2006). "Lenalidomide in the Myelodysplastic Syndrome with Chromosome 5q Deletion". New England Journal of Medicine. 355 (14): 1456–1465. doi:10.1056/NEJMoa061292. ISSN 0028-4793.
- ↑ Ebert, Benjamin L.; et al. (2008). "Identification of RPS14 as a 5q- syndrome gene by RNA interference screen". Nature. 451 (7176): 335–339. doi:10.1038/nature06494. ISSN 0028-0836. PMC 3771855. PMID 18202658.CS1 maint: PMC format (link)
- ↑ 13.0 13.1 Schneider, Rebekka K.; et al. (2014). "Role of Casein Kinase 1A1 in the Biology and Targeted Therapy of del(5q) MDS". Cancer Cell. 26 (4): 509–520. doi:10.1016/j.ccr.2014.08.001. PMC 4199102. PMID 25242043.CS1 maint: PMC format (link)
- ↑ Starczynowski, Daniel T; et al. (2010). "Identification of miR-145 and miR-146a as mediators of the 5q– syndrome phenotype". Nature Medicine. 16 (1): 49–58. doi:10.1038/nm.2054. ISSN 1078-8956.
- ↑ Patnaik, M M; et al. (2010). "WHO-defined 'myelodysplastic syndrome with isolated del(5q)' in 88 consecutive patients: survival data, leukemic transformation rates and prevalence of JAK2, MPL and IDH mutations". Leukemia. 24 (7): 1283–1289. doi:10.1038/leu.2010.105. ISSN 0887-6924. PMC 3035970. PMID 20485371.CS1 maint: PMC format (link)
- ↑ Heuser, Michael; et al. (2018). "Epigenetics in myelodysplastic syndromes". Seminars in Cancer Biology. 51: 170–179. doi:10.1016/j.semcancer.2017.07.009.
- ↑ Barlow, Jillian L; et al. (2010). "A p53-dependent mechanism underlies macrocytic anemia in a mouse model of human 5q– syndrome". Nature Medicine. 16 (1): 59–66. doi:10.1038/nm.2063. ISSN 1078-8956. PMC 2803774. PMID 19966810.CS1 maint: PMC format (link)
- ↑ Starczynowski, Daniel T; et al. (2010). "Identification of miR-145 and miR-146a as mediators of the 5q– syndrome phenotype". Nature Medicine. 16 (1): 49–58. doi:10.1038/nm.2054. ISSN 1078-8956.
Notes
*Primary authors will typically be those that initially create and complete the content of a page. If a subsequent user modifies the content and feels the effort put forth is of high enough significance to warrant listing in the authorship section, please contact the CCGA coordinators (contact information provided on the homepage). Additional global feedback or concerns are also welcome. *Citation of this Page: “Myelodysplastic neoplasm with low blasts and 5q deletion”. Compendium of Cancer Genome Aberrations (CCGA), Cancer Genomics Consortium (CGC), updated 09/6/2024, https://ccga.io/index.php/HAEM5:Myelodysplastic_neoplasm_with_low_blasts_and_5q_deletion.